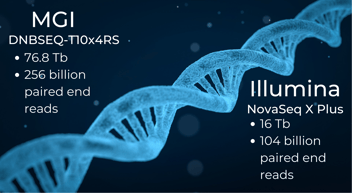
There are several sensitive methods for investigating gene expression on a large scale.
Microarrays are widely used for rapid, high-throughput screening of differential gene expression for biological, medical and clinical research and the collective data from these studies is available in several public databases (e.g., EMBL-EBI Expression Atlas, GTex and Gene Expression Omnibus).
Microarrays rely upon the direct hybridization of specific probes to known target RNA molecules in the sample and the results are provided as a numerical count of each RNA-bound probe.
While these methods are powerful tools for studying the response to treatment of a cellular model in terms of known RNA targets, they offer little opportunity for discovery of novel transcripts or the unanticipated but potentially important changes in the expression of noncoding transcripts.
RNA-seq, in contrast, offers the opportunity to interrogate the entire transcriptome at a given moment at single-base resolution without prior knowledge of genes that might be impacted by a particular treatment or condition, thus providing a more comprehensive, hypothesis-free view of the transcriptome.
These benefits are clearly shown by
Rao et al., in which data obtained by RNA-seq and microarrays were compared in a toxicogenomic study in rats: compared to microarrays, RNA-seq data yielded additional differentially expressed genes (DEGs) that enriched their knowledge of key pathways affected by test compounds, as well as additional relevant pathways.
These results clearly indicate that RNA-Seq is therefore likely to generate more insight into mechanisms of toxicity compared to microarrays thanks to its wider dynamic range, greater sensitivity and ability to identify a larger number of transcripts.
Now, the main challenge to a full deployment of RNA-seq in the drug discovery field remains the need for cost-efficient and high-throughput workflows, which make it feasible – economically and practically – to perform RNA-seq on tens of thousands of RNA samples.
Please
contact us to know more about the existing RNA-seq solutions designed for drug screening.